AI models and systems for data-driven scientific discovery.
Foundation models trained on multi-omic experimental data, not just language. Query our models or train your own.
Trusted by researchers at










Archimedes Foundation Model
AI Foundation Models for Biology
We train AI models on real multi-omic experimental data — not just published text. The result: foundation models that understand biological systems from the data up.
247K+
nodes
165M+
relationships
167M+
parameters
30+
different tissues
Use our models
Query Archimedes through ARC to generate hypotheses, find hidden connections, and analyze your data against our biological knowledge graph.
Train your own
Fine-tune on your lab’s data or build fully custom models. Your proprietary datasets stay private while benefiting from our architecture and training infrastructure. Your models, your data, your discoveries.
All your research, one place
- Organize experiments, datasets, and analyses by research question
- Local and cloud storage with workspaces
- Agentic search across all your data and projects
Curate your paper library
- Import by DOI — metadata and abstract auto-extracted
- Read and annotate PDFs inline
- AI summaries, methods extraction, and linking to your projects
See how your ideas connect
- Notes with bidirectional links
- Visual knowledge graph across your research
- AI digests surface patterns and connect ideas you might miss
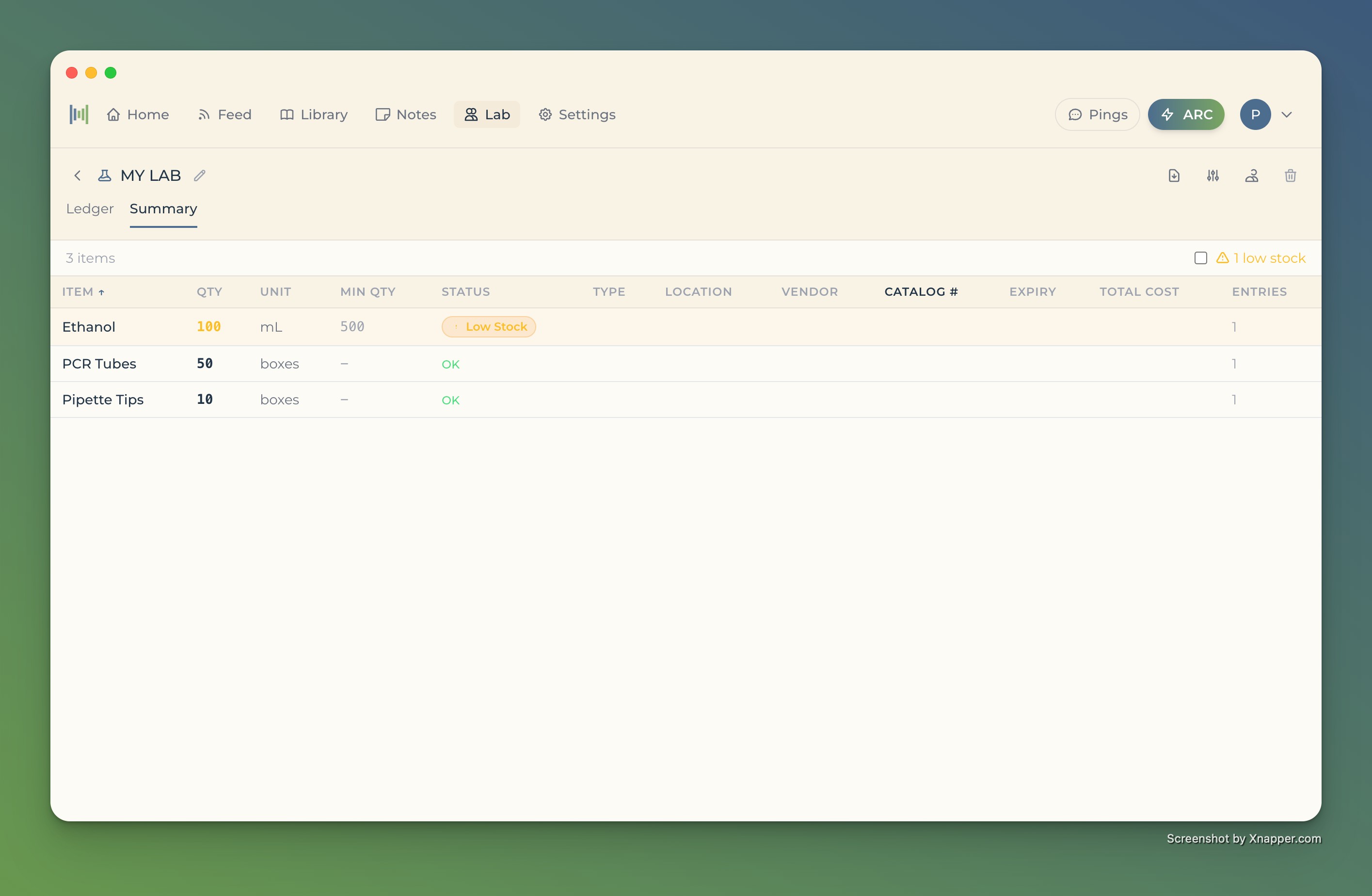
Track everything in your lab
- Reagents, consumables, and equipment in one view
- Automated low stock alerts before you run out
- Less time on inventory, more time at the bench
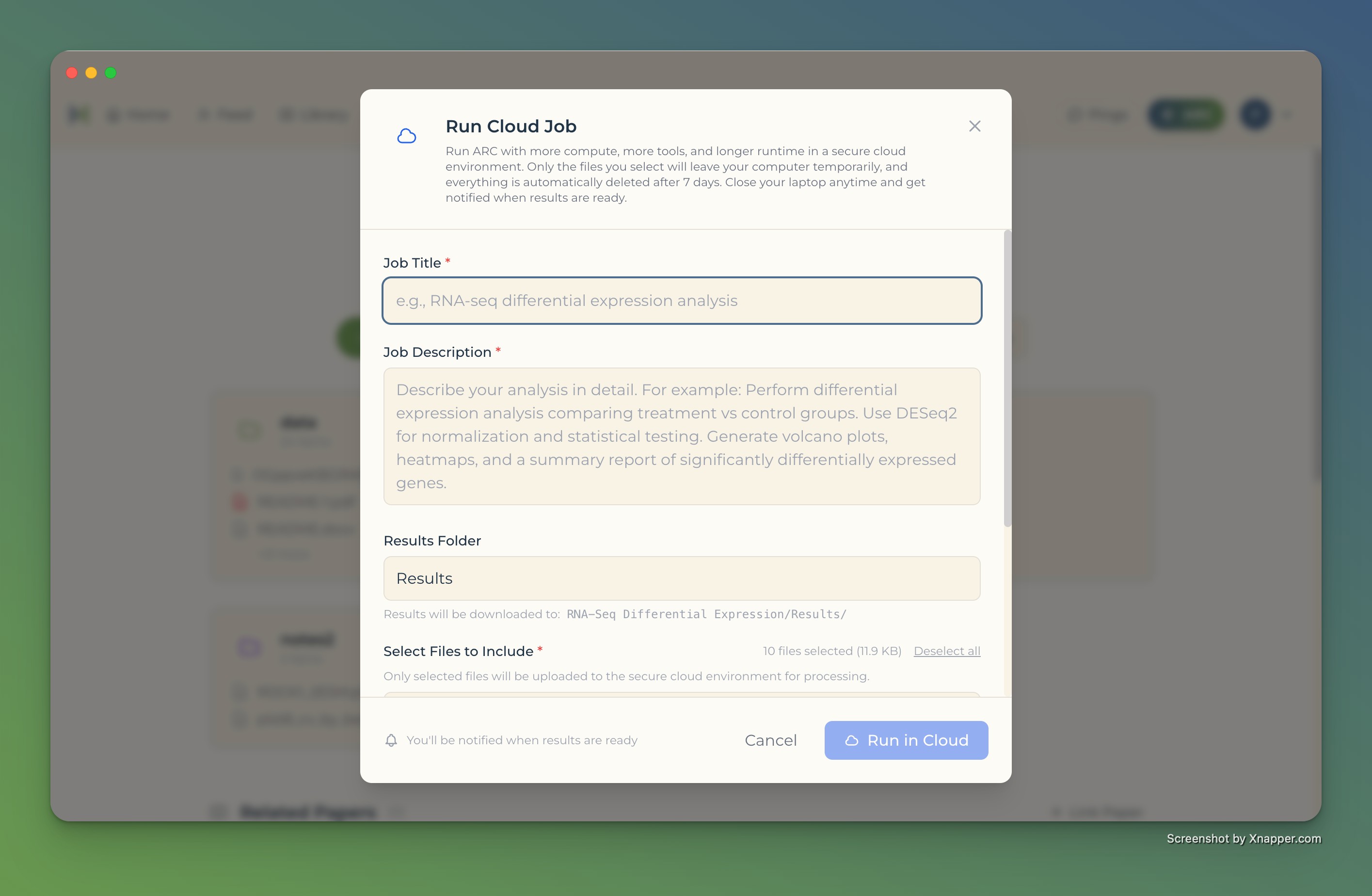
Cloud compute, from your desktop
- Run heavy analyses without leaving Bench
- Select data and define the job
- Results sync back to your project
Get your sequencing data, ready to analyze
- Transfer data from FTP/SFTP servers directly to Heureka Cloud
- Auto-detect FASTQ samples — supports 10x Illumina, SRA, and generic naming
- Run scRNA-seq or metagenomics pipelines, results delivered to your project

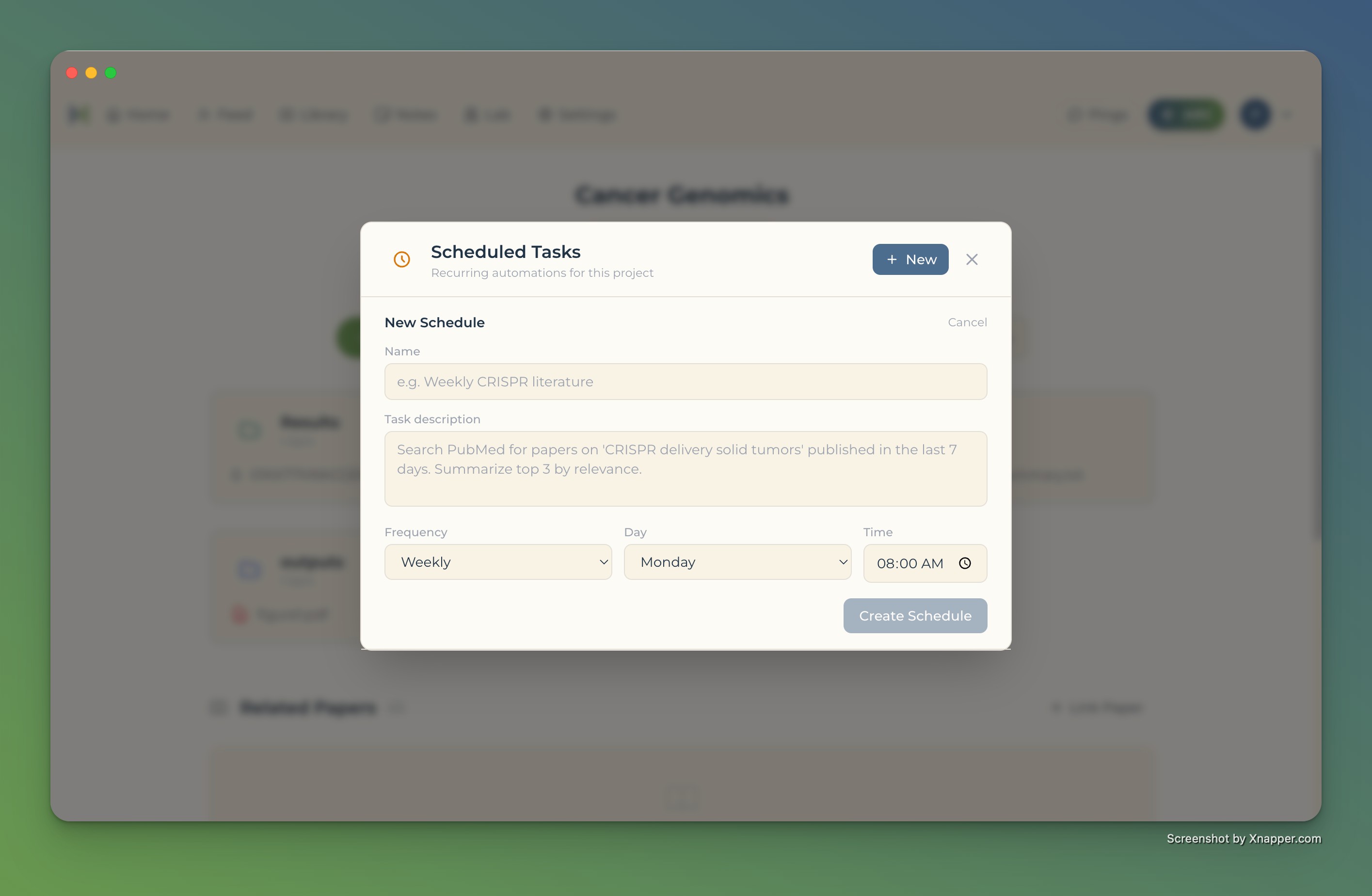
Scheduled tasks that run themselves
- Define what to analyze — ARC handles the rest
- Literature scans, hypothesis generation, data analysis on schedule
- Results delivered daily, weekly, or custom
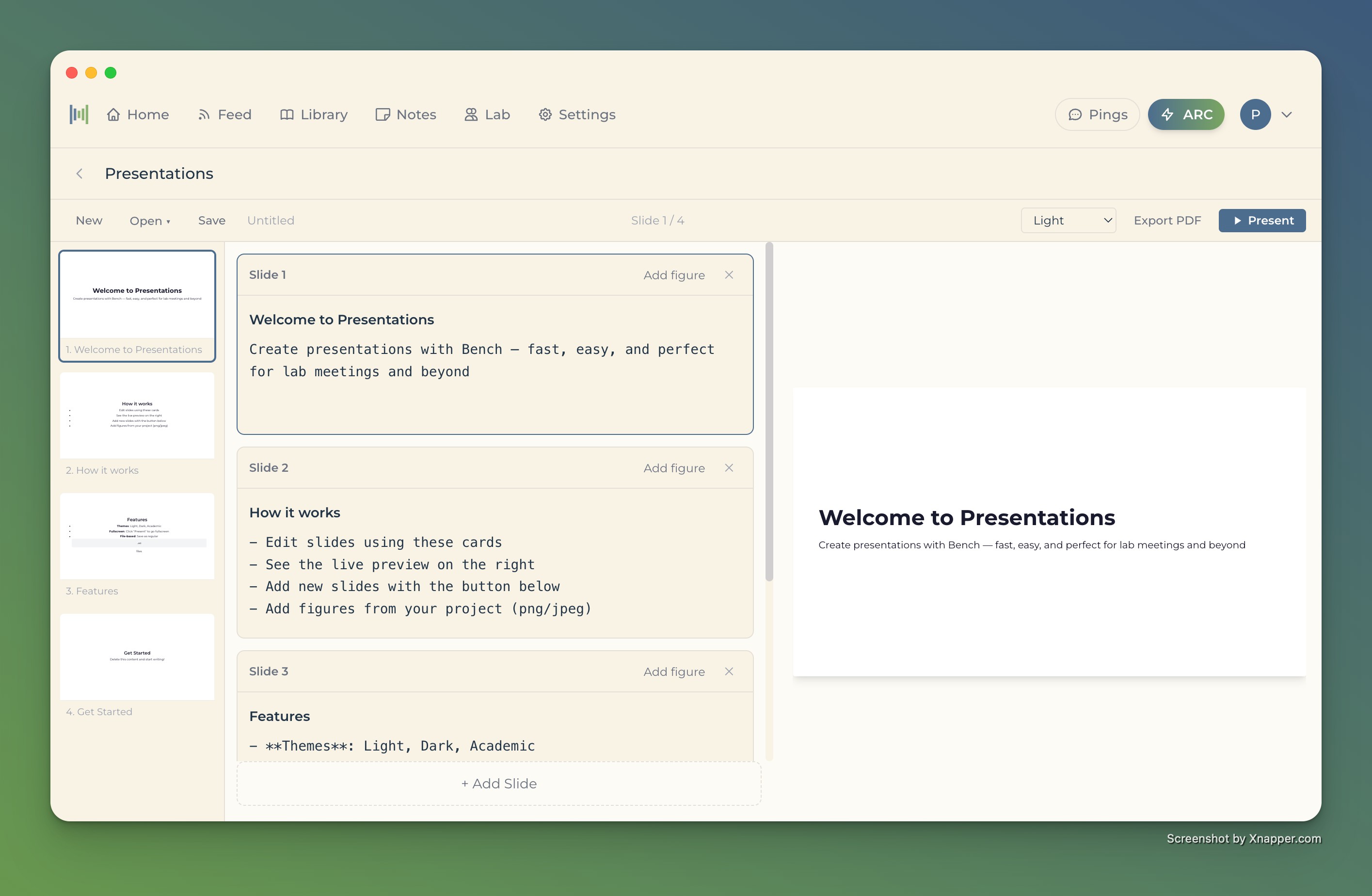
From findings to presentation
- Build slide decks from your research
- Slides auto-update when your underlying data changes
- Less time formatting, more time on your research
Published Research
Work Built with Heureka
"Our pathway coessentiality analysis identified specific metabolic pathways linked to complex II, including purine nucleotides. Among ~16,000 genes, loss of nucleotide biosynthesis enzymes strongly sensitized AML cells to complex II inhibition. In a syngeneic AML mouse model, targeting complex II leads to rapid disease regression and extends survival. These findings establish complex II as a central regulator of de novo purine biosynthesis and a promising therapeutic target in AML."

"When we don't use AI, we understand maybe 10% of the message, and 90% of the patterns go unidentified. AI helps us to do that. It helps us identify mechanisms of action... It allows us to fine-tune our approach in terms of interaction with other genes. It also helps us identify the patients most likely to respond... AI is not an advantage for biotech, it's a necessity."

Download Heureka Bench
No subscription. Your data stays on your machine.
